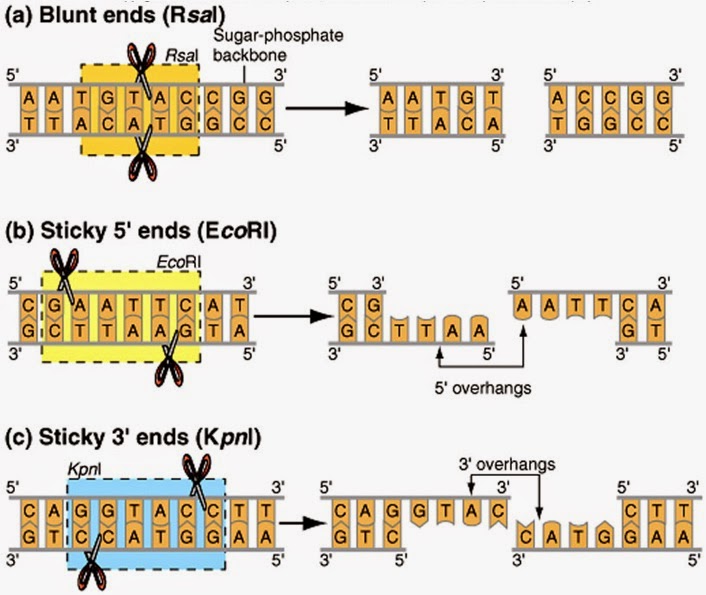
The length and the sequence of the recognition site varies from enzyme to enzyme, so that on average, enzymes with shorter recognition sites cleave DNA at more places, and produce smaller fragments than enzymes with longer recognition sites. There are presently hundreds of commercially available restriction enzymes, each with unique recognition sequences which allow the isolation of virtually any region of the genome. There is no limit to the design of such DNA rearrangements, as long as there are appropriate restriction sites in the DNA sequences to be restructured. The backbone of the resulting recombinant DNA the linkage of DNA from a human and a bacterial cell is indistinguishable from the original uncut DNA. A simple enzymatic reaction reforms the continuous double-stranded DNA across the breaks with a smooth splice. For example, a human DNA restriction fragment can be fused with the sticky end of bacterial DNA that was cut with the same enzyme. The sticky ends of a fragment of human DNA can then be joined to complementary ends of DNA cut from another source. Cleavage of DNA from any organism with a particular restriction enzyme always generates the same complementary ends on either side of the cut. The ragged or "sticky" ends that remain after restriction enzyme cleavage are actually a key aspect of recombinant DNA technology. At any given enzyme recognition cleavage site, these overhangs have unique and complementary sequences. Most restriction enzymes cleave their palindromic recognition sequences asymmetrically, leaving a single-stranded overhang at each side of the cut (see Figure 4, top). This allows each half of the restriction enzyme scissors to recognize the same sequence on both DNA strands. coli recognizes and cuts the sequence GAATTC in double-strand DNA at the GA and AG junctions. These recognition sites, which occur randomly along the DNA of any organism, consist of a short symmetrical sequence motif, or palindrome, which is repeated in opposing orientations on both strands of the DNA double helix. The restriction enzyme synthesized by specific species of bacteria recognizes a specific nucleotide sequence. The discovery that bacteria contain enzymes, called restriction endonucleases, that act like molecular scissors, cutting the DNA backbone at predictable sequence motifs, was a major breakthrough in the manipulation of DNA.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
